It was termed the “TATA box” as it contains a consensus sequence characterized by repeating T and A base pairs. The TATA box is considered a non-coding DNA sequence (also known as a cis-regulatory element). 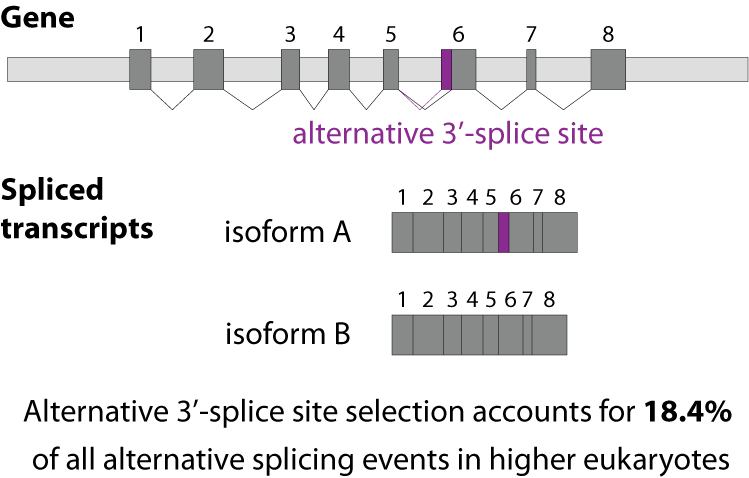
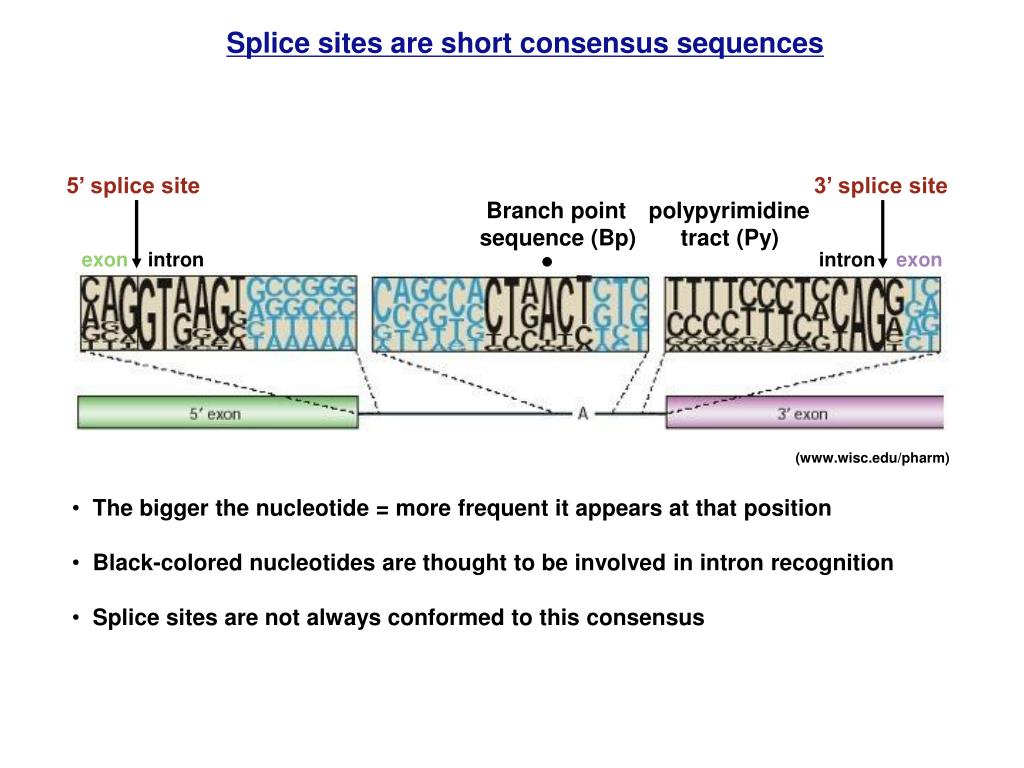
Often conserved sequences reflect a common function or binding domain. How do you identify a consensus sequence?Ī consensus sequence is determined by aligning many nucleotide (or protein) sequences that share a common function, then determining the most commonly expressed nucleotide (or amino acid) at each position. The highly conserved base at the branch point site is A whereas the 3′ splice site is AG rich and the 5′ splice site is GU rich. How many consensus sequences for splicing are found in an exon? Explanation: None of the consensus sequences for splicing are found in an exon. How many consensus sequences for splicing are found in an exon? Mutations, including splice site mutations, in the underlying DNA or errors during transcription can activate a cryptic splice site in part of the transcript that usually is not spliced. Several BP-dedicated bioinformatics tools, including HSF, SVM-BPfinder, BPP, Branchpointer, LaBranchoR and RNABPS were developed during the last decade. The canonical splice sites are those originally described and most commonly found (like in ~99% of introns) and have GT at the donor site (just after the 5′ end of the cut) and AG at the acceptor site (just before the 3′ end of the cut).Ī cryptic splice site is a mRNA sequence that has the potential for interacting with the spliceosome. Background Branch points (BPs) map within short motifs upstream of acceptor splice sites (3’ss) and are essential for splicing of pre-mature mRNA. The phrase also refers to an actual sequence which approximates the theoretical consensus. Most commonly, the RNA sequence that is removed begins with the dinucleotide GU at its 5′ end, and ends with AG at its 3′ end.Ī theoretical representative nucleotide or amino acid sequence in which each nucleotide or amino acid is the one which occurs most frequently at that site in the different sequences which occur in nature. These sites are found at the 5′ and 3′ ends of introns. Introns are removed from primary transcripts by cleavage at conserved sequences called splice sites. What are the consensus sequences seen at 5 and 3 ends of an intron? In humans, we identified 42 different consensus sequences that are each present in at least 100 human introns. How many consensus sequences are required to splice an intron? Little is known about the interactions formed by these three nucleotides in the spliceosome. The YAG/ consensus sequence at the 3′ end of introns (the slash indicates the location of the 3′ splice site) is essential for catalysis of the second step of pre-mRNA splicing. What is the 3 splice site consensus?Ībstract. These consensus sequences include nearly invariant dinucleotides at each end of the intron, GT at the 5′ end of the intron, and AG at the 3′ end of the intron. Splice Site Consensus (jump to matrices) It is well-established that nearly all splice sites conform to consensus sequences (matrices).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
